Computational analysis and prediction of PE_PGRS proteins using machine learning - Computational and Structural Biotechnology Journal
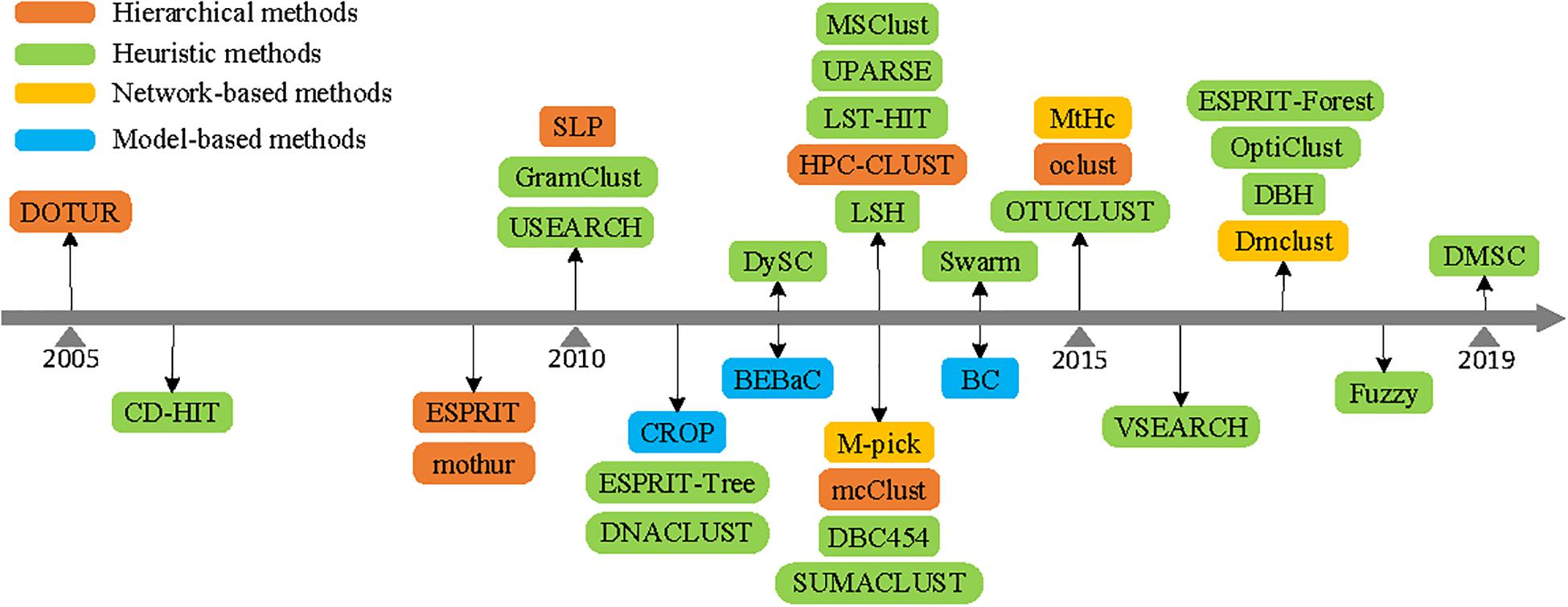
Frontiers | Comparison of Methods for Picking the Operational Taxonomic Units From Amplicon Sequences

kClust: fast and sensitive clustering of large protein sequence databases | BMC Bioinformatics | Full Text
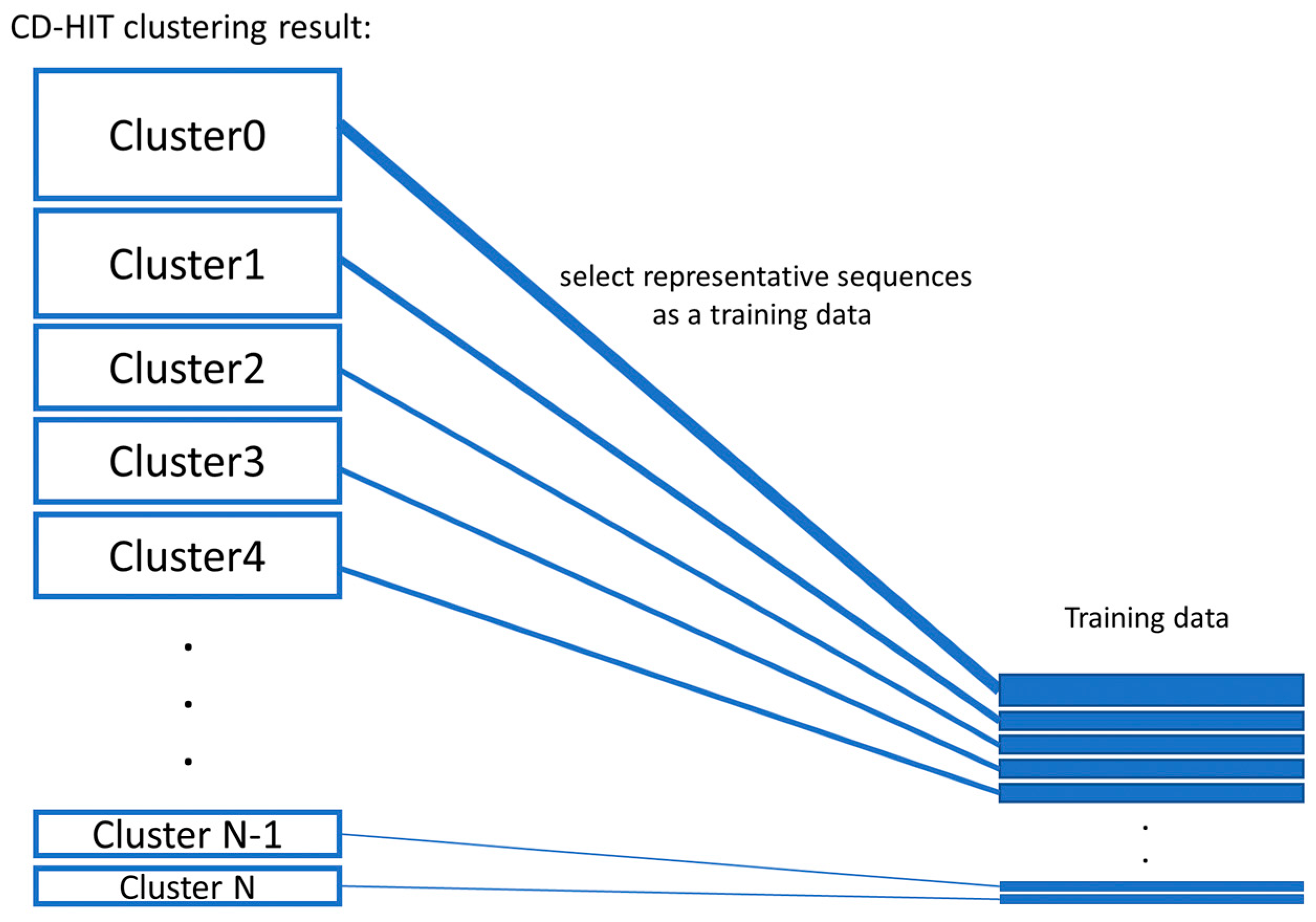
Genes | Free Full-Text | Ensemble-AMPPred: Robust AMP Prediction and Recognition Using the Ensemble Learning Method with a New Hybrid Feature for Differentiating AMPs

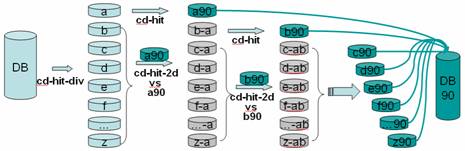
Fast Program for Clustering and Comparing Large Sets of Protein or Nucleotide Sequences | SpringerLink

Clustering biological sequences with dynamic sequence similarity threshold | BMC Bioinformatics | Full Text
![PDF] Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences | Semantic Scholar PDF] Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/f77ee5140d2c9408165c6cb756344fa6b0067f10/2-Table1-1.png)
PDF] Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences | Semantic Scholar

An example illustrating how the CD-HIT main paradigm works. Record 1 is... | Download Scientific Diagram

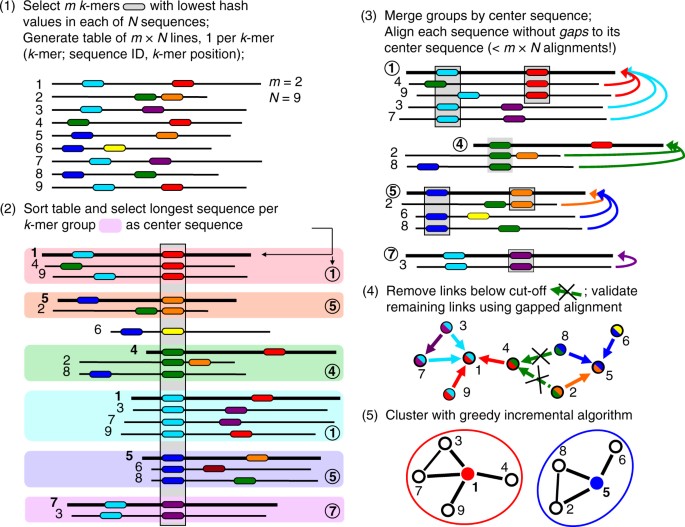






![Swarm: robust and fast clustering method for amplicon-based studies [PeerJ] Swarm: robust and fast clustering method for amplicon-based studies [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2014/593/1/fig-2-full.png)

