Discover the new Expasy.org, the Swiss Bioinformatics Resource Portal | SIB Swiss Institute of Bioinformatics

SIB on X: "“Whether you are a #PhDStudent looking for the ideal tool to analyse your #transcriptomics #data, or teaching undergraduates about #bioinformatics: #Expasy is the gateway to finding answers.” Christine Durinx,

Physiochemical Characterization and Domain Annotation of ORF1ab Polyprotein of Novel Corona Virus 19 | SpringerLink

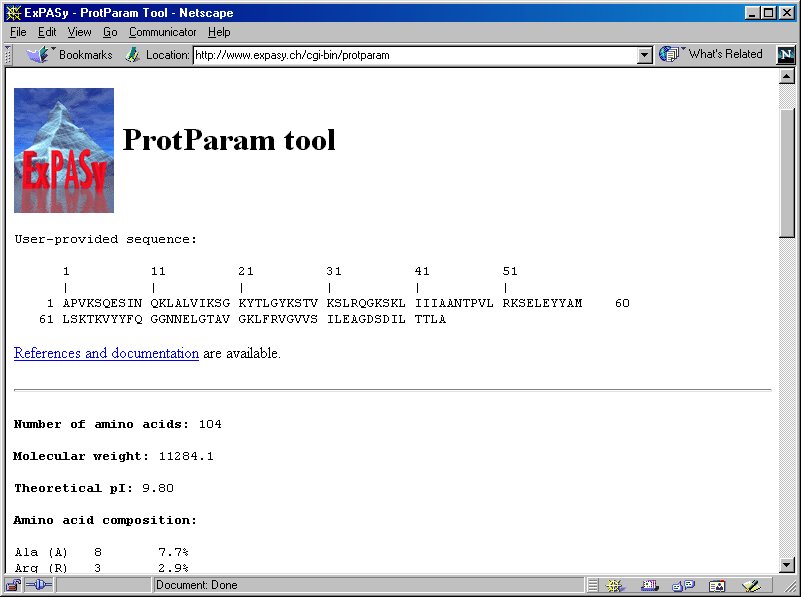
SOLVED: Use EXPASY Protparam tool to calculate the molecular weight, number of amino acids, and the theoretical pI, extinction coefficient under fully reducing conditions, and expected absorbance for 1 mg/ml protein. Use

Protparam tool for prediction of physical and chemical properties of proteins | Protparam tutorial - YouTube