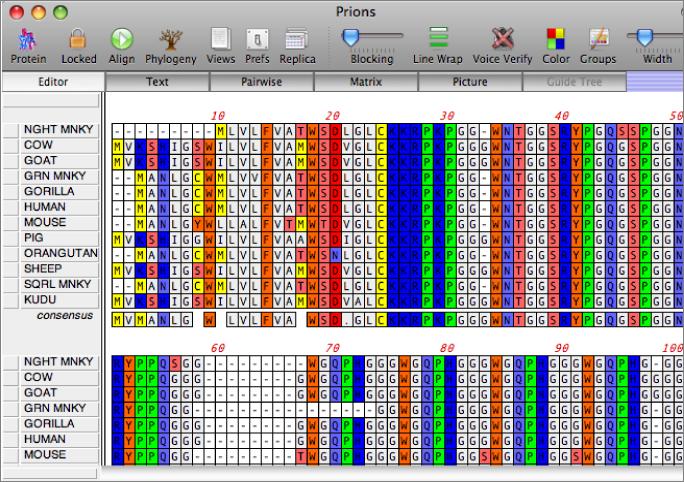
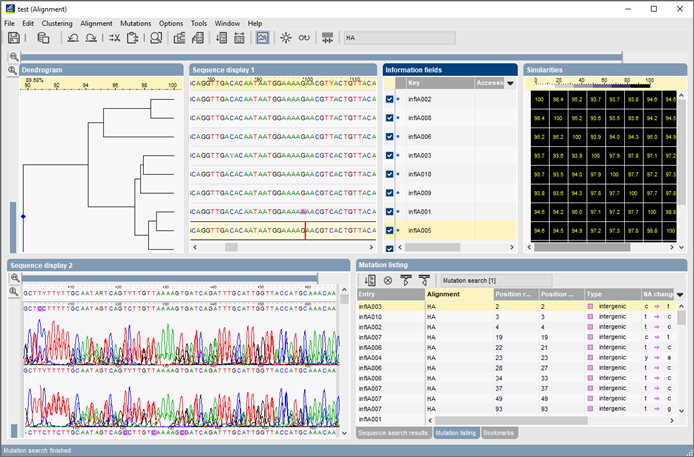
Figure 2 from The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. | Semantic Scholar
Software and Databases | The Barton Group | Bioinformatics Research | Division of Computational Biology | University of Dundee

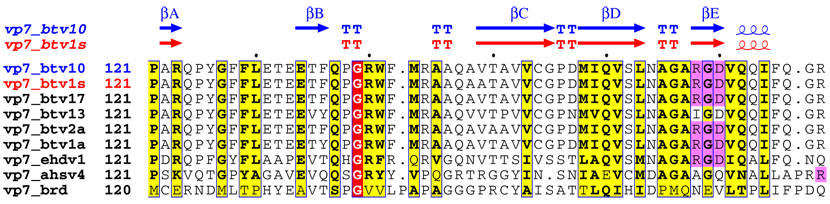
Multiple sequence alignment using ClustalW software. The Q33 residue is... | Download Scientific Diagram